Both this repository and its sister, Mathematica repository pareto_interactions, supplement our preprint: Emergence of division of labor in tissues through cell interactions and spatial cues.
In our work, we:-
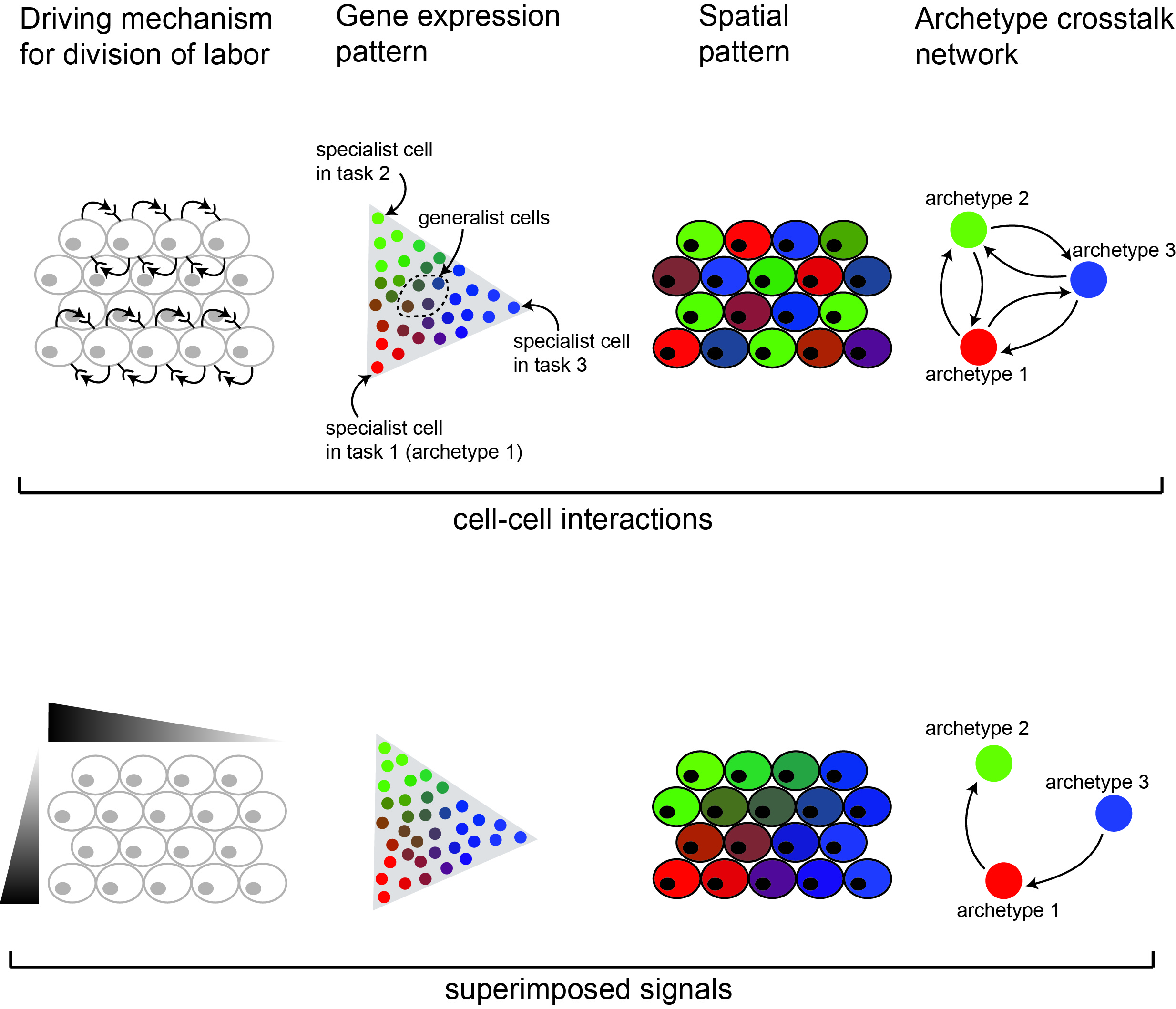
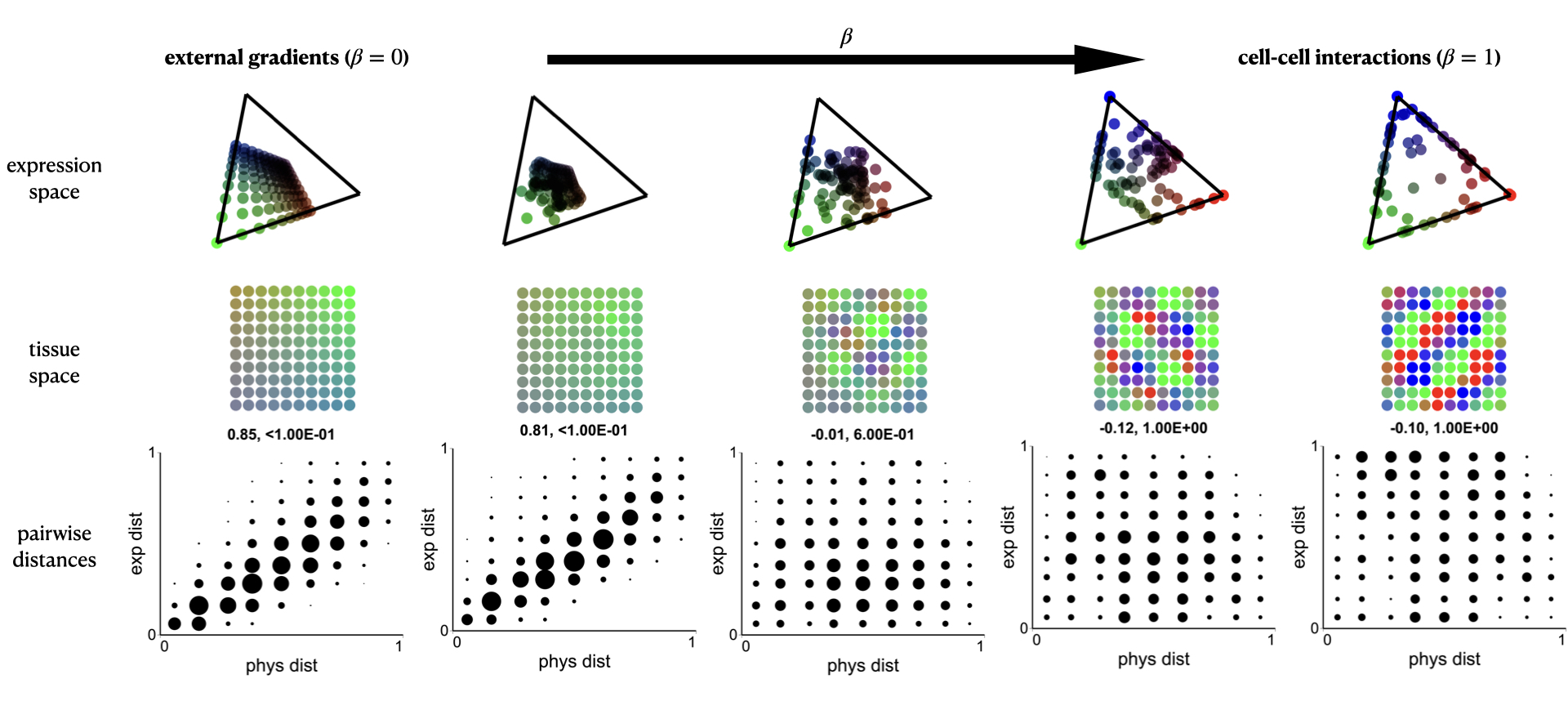
develop a computational framework to investigate the contribution of global gradients across the tissue and of self-organization mechanisms towards the division of labor within a cell-type population.
-
identify distinguishable expression patterns emerging from cell-cell interactions vs. instructive signals.
-
propose a method to construct ligand-receptor networks between specialist cells and use it to infer division-of-labor mechanisms from single-cell RNA-seq and spatial transcriptomics data of stromal, epithelial, and immune cells.
This repository contains:
- Simulated optimization of Pareto-optimal task and spatial distributions.
- Plotting and statistical significance testing of task distances verses physical distances.
- Analysis of colon fibroblast Slide-seq data (Avraham-Davidi et al.).
- Projection and analysis of colon fibroblast single-cell RNA-seq expression (Muhl et al.) onto Slide-seq data (Avraham-Davidi et al.).
Create a clean environment with conda using the environment.yml file from the this repository:
conda env create -f "https://raw.githubusercontent.com/nitzanlab/pareto_interactions_py/master/environment.yml"
(For older versions of conda one needs to download the environment.yml and use the local file for installation.)
For install in an already existing environment, use pip of the latest release from github:
pip install pareto_interactions_py@git+https://github.com/nitzanlab/pareto_interactions_py.git
Feel free to contact us by mail.